  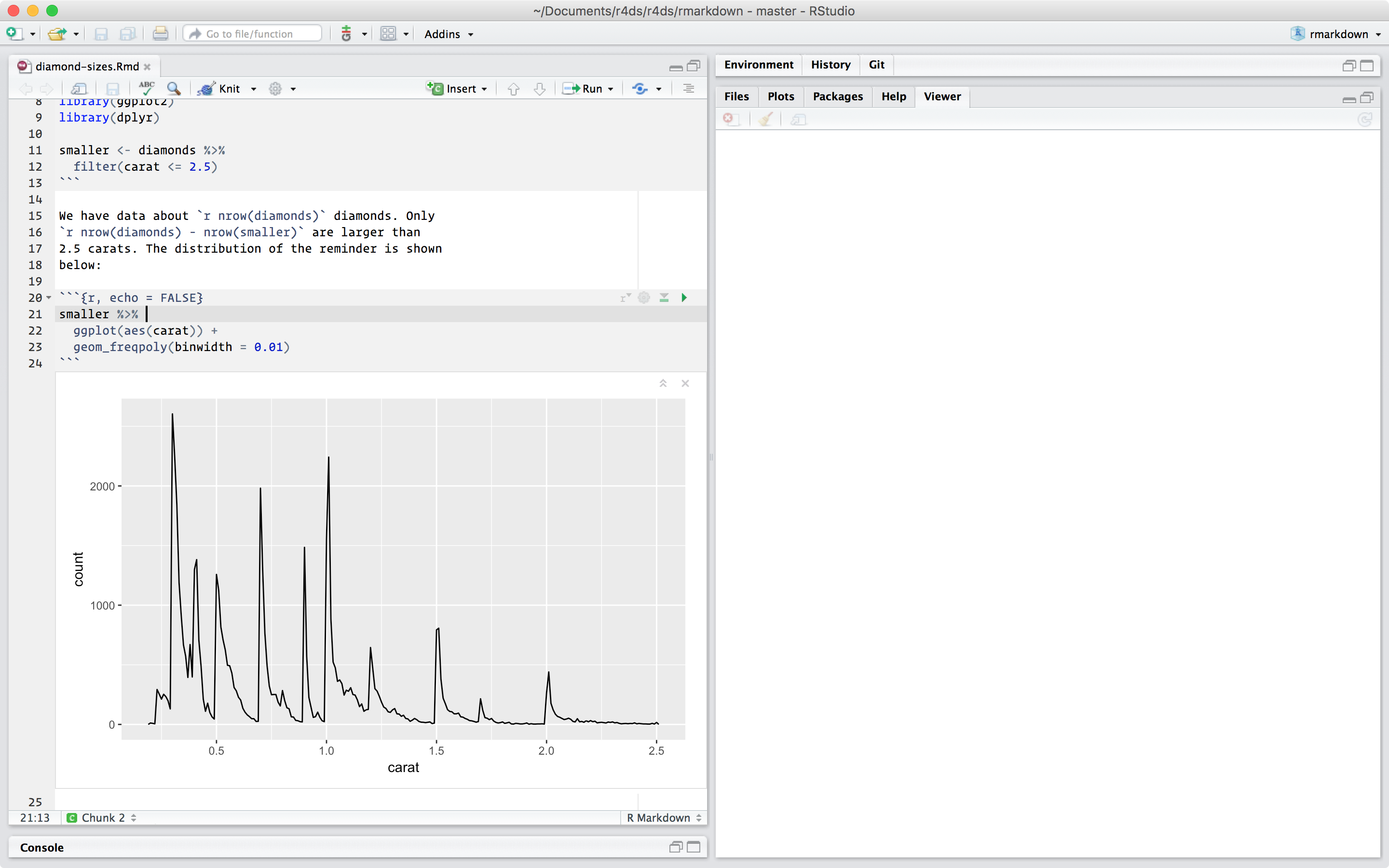
You can specify a code chunk by starting it ``` and ending it with ```. The template R Markdown script includes three code chunks. Text and code blocks are also included, and these will be explained in more detail below. 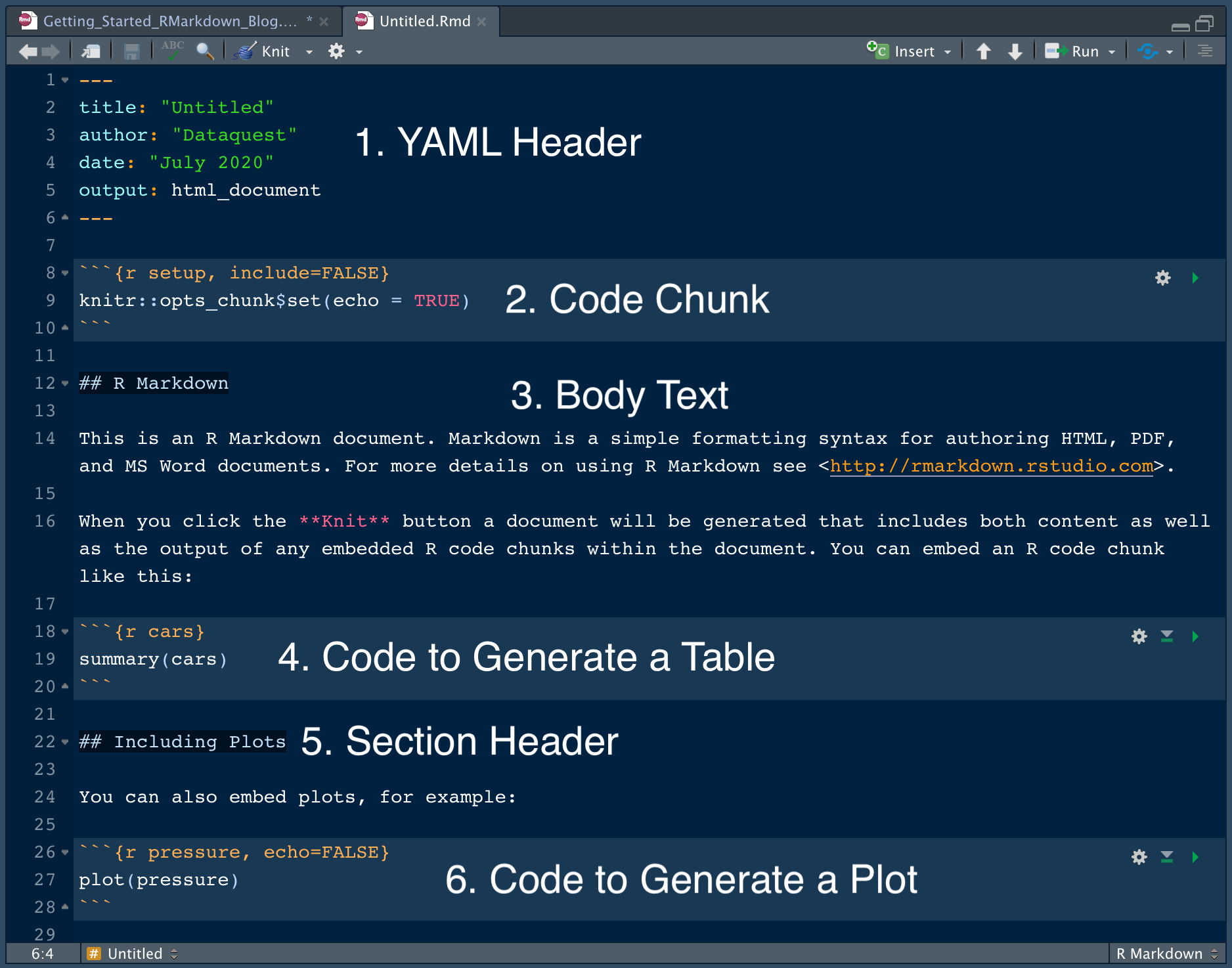
This is used by knitr during rendering to produce the correct file format. More setup options can be added if needed. This includes the set up information at the top of the page in between two lines of three dashes. A template R Markdown script is provided. The new R Markdown file should now have opened on the left hand side, above the console window. Give the document an appropriate name and choose “HTML” as the output format: You can also see from the left hand tab that R Markdown can be used to make other things besides documents, including presentations and shiny apps. All of these elements can be changed later if you change your mind. The following window then asks you for a title, the author and what format you’d like the final rendered file to be. To do this go, to the “Open” symbol in the top left hand corner and select “R Markdown” You should now be able to open a new R Markdown file. To do this, in the console panel (on the left), run: Once RStudio is installed and running, your window should look something like this:Īs well as installing RStudio, you’ll need to have the package for rmarkdown installed. For more details on this, see our workshop on Version Control with RStudio and Github. It allows you to view scripts, the console, your environment and plots all within one window. RStudio is an easy to use graphical user interface for running R. If you don’t already have RStudio, you can download it from the following link. 
The benefits of R Markdown are best appreciated when using it within RStudio. This video gives a great, short explanation of R Markdown. R Markdown is a variation on Markdown allowing it to be implemented in R. Due to it’s basic nature, you need none to very little programming knowledge in order to write in Markdown! You can see the original Markdown code here. This webpage has been written in Markdown and then github has rendered this to allow you to view it as a webpage. It was originally designed for web developers to allow for editing of web pages with an easy-to-read and easy-to-write plain text format. Markdown is a coding language that allows for text-to-HTML conversion. This tutorial has been largely inspired by the fantastic resources available at the R Markdown Website. It also has the ability to the render the R Markdown into easy-to-read documents including PDF, html or word document formats, allowing for easy production of reports. R Markdown is a nice solution to this situation, allowing you to group your code into “chunks” as well as acting like a notebook, with plots pictured directly below the code. Even with thorough notation of the script, this can often still be confusing. However, I’ve often found myself lost in a 1000 line script, trying to work out what each line of script is doing and what plots are being produced. As a researcher who uses R on a daily basis, I started out using R Scripts to record my research. R has several nice ways to record your activities, and to make these as reproducible as possible including R Scripts and R Markdown. However when it comes to statistics and plots, people are less cautious about recording what they have done. This is a common practice within the wet lab with all researchers keeping a lab book. It’s important during research to keep a thorough record of your analysis. Contact: Research field: Bioinformatician working on analysis of Next Generation Sequencing Data. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
